Serviços Personalizados
Journal
Artigo
Indicadores
-
 Citado por SciELO
Citado por SciELO -
 Acessos
Acessos
Links relacionados
-
 Similares em
SciELO
Similares em
SciELO
Compartilhar
Ciência e Técnica Vitivinícola
versão impressa ISSN 0254-0223
Ciência Téc. Vitiv. v.25 n.1 Dois Portos 2010
The portuguese Vitis vinifera L. germplasm: genetic relations between wild and cultivated vines
Jorge Cunha 1,2, Margarida Teixeira-Santos3, Manuela Veloso3, Luís Carneiro3, José Eiras-Dias1, Pedro Fevereiro2,4*
1) INRB, INIA, Dois Portos, Quinta da Almoinha, 2565-191 Dois Portos (Portugal).
2) ITQB, Universidade Nova de Lisboa, Av. da República, Apt. 127, 2781-901 Oeiras (Portugal).
3) INRB, INIA, Oeiras, Av. da República Quinta do Marquês, 2784-505 Oeiras (Portugal).
4) Universidade de Lisboa, Fac. Ciências, Dept. Biologia Vegetal, Campo Grande, 1749-016 Lisboa (Portugal).
ABSTRACT
Portuguese wild vine populations are in an apparent geographic fringe of the species distribution. Despite, Portugal offers a unique richness in autochthonous cultivated varieties that contributes to the overall diversity of worldwide grapevine. In the different Portuguese agro-ecosystems, grapevine plays an important role either as a border culture or an extensive crop. During the last years wild vine populations have been identified but only in southern riverside ecosystems. To conclude if the local wild vine germplasm is involved in the origin of cultivated grapevine we used the OIV recommend six nuclear and four chloroplastidial microsatellite loci to genotype and to established phylogenetic relationships between wild plants and cultivated grapevines. Both native sylvestris and vinifera subspecies have a high genetic diversity and a sizeable number of rare alleles. Portuguese wild vine populations showed a high level of intra-population diversity with most of the genetic diversity conserved within each population. Wild vines seem to form a continuum and there is no clear population division. Despite, a low, but still significant, genetic differentiation can be detected among the analyzed populations. There seem to be a close genetic relation between the wild plants and cultivated varieties. Both subspecies mostly share the A and B chlorotypes, tipical of the Iberian germplasms. In some cases we found a close genetic relation between cultivated varieties and wild plants. Finally, some native grapevine cultivars have a higher genetic diversity that reveals introgression of foreign gene pool. This study contributes to establish the range of existing genetic variability in the Portuguese native grapevine and wild vines germplasm. It also provides a baseline for future monitoring of the genetic diversity of the species in Portugal and contributes with data to construct a core collection to preserve the existing variability and delineate conservation strategies for the wild vines.
Key words: Vitis vinifera ssp. sylvestris; V. vinifera ssp. vinifera; nuclear SSR; Chloroplastidial SSR; multiple discriminant and principal coordinate analysis; genetic diversity.
O germoplasma português de Vitis vinifera L.: relações genéticas entre videiras selvagens e castas cultivadas
RESUMO
As populações portuguesas de videiras selvagens estão numa aparente orla da distribuição geográfica da espécie. No entanto Portugal apresenta uma riqueza única em castas autóctones, as quais contribuem para a diversidade global da vinha. A vinha tem um papel importante quer marginalmente, quer como cultura extensiva, nos diferentes agro-sistemas portugueses. Durante os últimos anos foram identificadas populações de videiras selvagens mas só nos ecossistemas ripícolas do Sul do país. Para se concluir se o germoplasma selvagem local está envolvido na origem das castas portuguesas utilizámos seis loci de microsatélites nucleares e quatro cloroplastidiais para estabelecer relações filogenéticas entre as videiras selvagens e as castas cultivadas. Quer a subspécie sylvestris quer a viniferaapresentam uma diversidade genética elevada e um número apreciável de alelos raros. As populações portuguesas de videiras selvagens apresentam um nível elevado de variabilidade intra-populacional, estando a maior parte da diversidade genética conservada dentro de cada população. As videiras selvagens parecem formar um contínuo espacial e não existe uma clara divisão entre as populações. No entanto, é possível detectar uma baixa, mas mesmo assim significativa, diferenciação genética entre as populações analisadas. Parece existir uma estreita relação genética entre as videiras selvagens e as castas cultivadas. Ambas as subespécies partilham os mesmos clorótipos A e B, típicos do germoplasma Ibérico. Em alguns casos foram encontradas estreitas relações genéticas entre castas cultivadas e videiras selvagens. Finalmente algumas castas nativas têm uma maior diversidade genética, revelando uma introgressão de germoplasma estrangeiro. Este estudo contribui para definir a amplitude existente da diversidade genética do germoplasma das duas subspécies de Vitis presentes em Portugal. Também fornece uma linha de base para a futura monitorização da diversidade genética da espécie em Portugal e contribui com dados para a construção de uma colecção nuclear para preservar a variabilidade existente e para delinear estratégias de conservação para as vinhas selvagens.
Palavras-Chave: Vitis vinifera ssp. sylvestris; V. vinifera ssp. vinifera; SSR nucleares; SSR Cloroplastidiais; análise multivariada discriminante e em coordenadas principais, diversidade genética.
INTRODUCTION
The Eurasian Vitis vinifera L. comprise two subspecies, the wild vine ssp sylvestris (Gmelin) Hegi and the cultivated vine ssp vinifera L. Most botanists consider the wild ancestral grape as the primitive form of the cultivated grapevine due to the close morphological resemblance and free gene flow between them (Heywood and Zohary, 1991). Wild vine is a dioecious liana inhabiting flooded areas, on screens (colluvial sites) of hilly humid slopes and occasionally on coastal sheers and beaches from the South Atlantic coast of Europe to the Western Himalayas (Hegi, 1925). Todays reduced habitat of the subspecies is a result of the dramatic spread of pathogens from North America (philloxera, oidium and mildew) during the last 150 years, together with the fragmentation of its habitat due to intensive river management, forest cutting and removal of tutor trees (Grassi et al., 2006) thus reducing the overall diversity of this particular ecosystem. Even though it is in the IUCN list of endangered species since 1995, it has no special protection status in the majority of the co untries where the subspecies is found. Wild vine has been used by human since the early Neolithic as shown by the amount of pips recorded in prehistorically sites (Buxó, 2008) and until recently, at least in the Iberian Peninsula, was used to produce vinegar, enhance the colour of wine and prepare folk medicines (López Martínez et al., 2001).
Archaeological and historical information indicate the Near East or the Transcaucasian region as the centre for grapevine domestication (Rivera and Walker, 1989). Vitis vinifera spp vinifera is a worldwide fruit crop that has been cultivated for millennia for fresh and dried use as well as for wine production. Grapevine cultivation in the original range of the species played an important role in establishing particular agro-ecosystems adapted to local environment. Molecular analysis have revealed that cultivated grapevines in different regions harbour a high genetic diversity and heterozygosity suggesting also that secondary domestication events have taken place with the contribution of local wild vine populations (Arroyo-Garcia et al., 2002). Domestication of grape involved a shift in the mode of reproduction from dioecious to hermaphrodite, ensuring self-pollination without the need for external pollen donor. Nevertheless all the grapevines cultivars are highly heterozygous (Jaillon et al., 2007) suggesting its origin from cross pollination. Traditionally, a single vineyard comprised a number of varieties, but in the last two centuries there was a shift to single variety vineyards sometimes limited just to one clone. Currently a great number of wines are made from an extremely limited number of cultivars and clones, partially because their agronomical traits and oenological techniques are better known, partially due to pure fashion. This trend contributes to reduce the accumulated diversity of cultivated grapevine. The efficient use of germplasm resources depends on the adequate knowledge of genetic variations and of genetic relationships between populations or genotypes (Wan et al., 2008).
Traditionally morphological descriptors were used to characterize cultivars until the advent of molecular markers. Presently these have been successfully used in a wide range of applications such as assessing genetic diversity (Sefc et al., 2000), for linkage mapping (Doligez et al., 2002), cultivar identification and pedigree studies (Schneider et al., 2001, Crespan 2004). Microsatellites (SSR) are used to characterize grapevine cultivars and wild vines (Sefc et al.,1999, 2000) and to carry out genetic diversity analyses (Aradhya et al., 2003). Usually six loci are sufficient for differentiating between genotypes (This et al., 2004), but closely related cultivars require larger number of pairs (Meredith et al., 1999). Sequence variation at the chloroplastidial loci is extensively used to assess phylogenetic relationships among plant taxa, based on their low rate of sequence evolution, the almost absent recombination and single parent inheritance (Vendramin et al., 1996). Chloroplastidial microsatellites (cpSSRs) have been used to study genetic relationships among grapevine cultivars (Imazio et al., 2006), wild vines (Grassi et al., 2006) and relations between both subspecies (Arroyo-Garcia et al., 2006, De Mattia et al., 2008).
Portuguese wild vine populations are in an apparent geographic fringe of the species distribution but the country richness in cultivar diversity (Almandim et al., 2007) and the importance in allele contribution to the overall diversity of grapevine (Le Cunff et al., 2008) tells another story. In the different Portuguese agro-ecosystems, grapevine plays an important role either as a border culture or an extensive crop. Despite the wide distribution of grapevine cultivation the wild vine populations identified to date are restricted to southern riverside ecosystems. The aim of this study is to determine how the local wild vine populations contributed to the cultivated grapevine diversity using the recommend six nuclear microsatellites loci for genotyping (OIV, 2007) and the four chloroplastidial microsatellite loci to determine the maternal relationships. The establishment of the genetic diversity and the phylogenetic relations between the two germplasm pools will contribute to decide the best strategies to preserve the Portuguese germplasm.
MATERIALS AND METHODS
Plant material
Wild Vines
Riverbanks are the present environment range of Vitis vinifera ssp. sylvestris in south-western Europe. The wild subspecies is dioecious as opposed to cultivars that are mainly hermaphrodites. Its propagation is either by seed or self rooting. To sample wild plants we choose morphological distinct male and female from each population. A population is a group of plants inhabiting a section of a water stream within 1 km of range and 10km apart from the next group of plants. The number of plants per population range from 5 to 50. From the twelve already identified populations we choose to sample four of them and collecting sixty plants. These four populations represent the range of environments so far identified in Portugal occupied by this subspecies. Population 01 (Montemor-o-Novo) is settled in the Almansor river margins (Tagus southern tributary) and population 02 (Castelo Branco) in the Pônsul river margins (Tagus northern tributary), both belonging to the hydrographical Tagus river basin. These populations are more than 100kms apart from each other. Population 01 is surrounded by commercial vineyards and is less than 70km from the Atlantic coast. Population 02 is located in a protected area (Parque Natural do Tejo international) 200km from the Tagus river mouth. Population 04 (Alcácer do Sal) is located in the Sado river plain, 20Km from the sea and surrounded by rice fields. Population 05 (Portel) is located in the Corte rivulet, a tributary of the Guadiana river, just near of the Alqueva dam. The location of all found populations is shown in figure 1.
Young leaves from the sixty Vitis vinifera ssp. sylvestris (Gmelin) Hegi plants were collected at bloom to identify the plant sex.
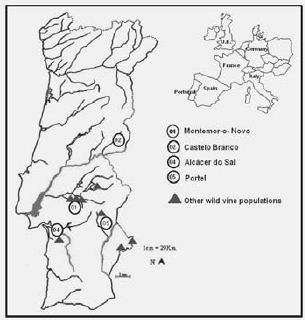
Figure 1 – Map of Portugal with the locations of the wild vine populations.
Mapa de Portugal com a localização das populações de videira selvagem
Grapevine cultivars
Seventy six grapevine cultivars (fifty seven native Portuguese and nineteen international) of Vitis vinifera ssp. vinifera were collected from the Portuguese National Ampelografic Collection (PRT 051), INIA Dois Portos, INRB I.P. (Quinta da Almoinha). The Portuguese cultivars were chosen among the ones traditionally planted in the wine areas adjacent to the wild vine populations. International cultivars were added for comparison with similar studies. Table I lists all the accessions used.
List of grapevine cultivars and wild vines genotyped
Lista das castas e das plantas selvagens genotipadas
Nuclear microsatellite genotyping
Total DNA was extracted and isolated from leaves as described by Almadanim et al. (2007) and amplified using six pairs of primers flanking nuclear SSR sequences, VVMD5, VVMD7, VVMD27, VrZag62, VrZag79 and VVS2 which are suggested by OIV for Vitis characterization (Table II). PCR reactions from nuclear SSRs were performed in 20 µl volume and the reaction mixture contained 10 ng DNA, 200 µM of each dNTP, 0.5U of Taq DNA polymerase(MB Fermentas), 2µl of 10× PCR buffer [(NH4)2SO4 – Fermentas], 2.5mM MgCl2), 0.3 µM of each primer except for VVS2 which uses only 0.125µM. The amplification parameters used were: initial denaturation at 95 ◦C for 5 min, followed by 35 cycles with a temperature profile of 95 ◦C for 20 s, 55 ◦C for 30 s and 72 ◦C for 30 s for all loci except for VVS2 which uses 50 ◦C for annealing temperature and a final extension time of 5 min at 72 ◦C. one cycle at 95 ºC for 5 min; 35 cycles at 95 ºC for 20s, annealing temperature for 30s, and 72 ºC for 30s; followed by 05min at 72 ºC Allelic size determination was carried out through capillary electrophoresis in a CEQ 8000 Genetic Analysis System (Beckman Coulter).
Source, locus denomination, repeat motifs, size ranges, accessions (n), number of alleles (Na), observed (Ho), expected heterozygosity (He) and Polymorphism Information Content (PIC) for six analyzed nuclear microsatellite
Origem, denominação do locus, motivo repetido, gama de tamanhos, número do acesso (n), número de alelos (Na), heterozigocidade observada (Ho), e esperada (He) e Informação do Conteúdo Polimórfico (PIC) para os seis microsatélites nucleares analisados.
Chloroplastidial Microsatellite genotyping
All the native Portuguese plants used in this study were previously genotyped by us (Cunha et al., 2009). Nineteen international grapevine cultivars (Portuguese synonyms in brackets): Alicante Bouschet; Farana/Damaschino (Alicante Branco), Tempranillo (Aragonez), Trouseaux (Bastardo), Mencia (Jaen), Malvasia di Lipari (Malvasia Candida), Muller-Thurgau, Riesling, Syrah, Teinturier, Coarna Neagra, Ahmer Bou Amer (Ferral), Aramon Noir, Cabernet Franc, Palomino Fino (Malvasia Rei), Sauvignon, Semillon, Trebbiano Toscano (Talia); and Verdello; were genotyped for the chloroplastidial SSR sequences, ccmp2, ccmp3, ccmp5 and ccmp10, using the pairs of primers suggested by Weising and Gardner (1999). Polymerase chain reactions (PCR) were carried out following the protocol described by Cunha et al. (2009). Each final amplification mix of 20 μl contained 10 ng DNA, 200 μM of each dNTP, 0.5U of Taq DNA polymerase (MB Fermentas), 2μl of 10× PCR buffer, 2.5mM MgCl2, 25ng of 33P-labeled primer, and 25ng of reverse primer. PCR conditions were: initial 5 min at 94ºC, followed by 35 cycles of 94°C for 45s, 52°C for 30s, and 72°C for 1min, with a final extension of 7min at 72ºC. Allelic sizes determination was carried out through capillary electrophoresis in a CEQ 8000 Genetic Analysis System (Beckman Coulter).
To confirm the amplified sequences, the PCR products were cloned into the Topo 4 vector (Invitrogen) and sequenced.
Data analysis
The MICROSAT program package (Minch et al., 1997) was used to calculate the proportion-of-shared-alleles distance between pairs of accessions to exclude identical nuclear genotypes. In all calculations, samples were considered as homozygous instead of heterozygous with a null allele when only one single allele was detected per locus.
The PowerMarker v3.23 program package software (Liu, 2002) was used to calculate the average number of alleles per locus (Na), the observed heterozygosity (Ho), the expected heterozygosity (He) or gene diversity and the Polymorphism Information Content (PIC) for each microsatellite locus as well as the average number of alleles Nal, Ho, He and inbreeding coefficient (f) in each population across all loci.
FSTAT v2.9.3.2 program package (Goudet, 1995) was used for calculating the allelic richness (Nar).
GenAlex6 program package (Peakall and Smouse, 2006) was used to assess the number of private alleles (Npr); to calculate the pair wise standard genetic distances (Nei, 1972) and the standard Fst (via Frequency) values; to calculate the molecular variance (AMOVA); and to determine the hierarchical distribution of genetic variance within populations and among populations. The significance of the f statistics was tested non-parametrically with 1000 permutations.
GENEPOP v3.4 program package (Raymond and Rousset, 1995) was used for testing genotypic frequencies in conformance to Hardy–Weinberg (HW) expectations, to test the loci for linkage disequilibrium and to estimate the significance of genotypic differentiation between population pairs. All probability tests were based on the Markov chain method (Rousset and Raymond, 1995) using 10.000 de-memorization steps, 100 batches and 5000 iterations per batch.
The NCSS 2000 program package (NCS, Kaysville, UT) was used for the multiple discriminant analysis, which first determines the average and the standard deviations of the variable of each group and them, determines the total correlation among variables, among and within the groups. Discriminant score, Eigenvalues, percentage and cumulative percentage due to each discriminant score, and the appropriate significant tests were also calculated.
The NTSYS-pc version 2.1 program package (Rohlf, 2000) was used for the Principal Coordinate Analysis. A minimum spanning tree is superimposed to represent the genetic relation among individual plants. To make possible the use of the chloroplastidial genotype data in this analysis we assumed it to be diploid and homozygous.
RESULTS
Nuclear Microsatellite diversity
When genotyped, seven wild vine accessions revealed to be clones, probably due to self rooting, so only the remaining fifty three were further analysed. A total of fifty three alleles were scored across the six nuclear loci, the number of alleles ranging from seven (VRZag62) to eleven (VVS2) with a mean value of 8.8 alleles per locus. For all loci most allele sizes varied in steps of more than two nucleotides and only locus VVS2 showed alleles that vary by steps of two. Allelic frequency ranged from 0.009 (alleles present in a single accession) to 0.750. For a reduced number of locus allelic frequencies are greater than 0.3 (Table III – in bold). The observed heterozygosity ranged from Ho=0.509 to Ho=0.736, with a mean value of 0.601. The expected heterozygosity was similar to the observed heterozygosity, ranging from He=0.588 to He=0.801, with a mean value of 0.676. All nuclear microsatellite loci scored were polymorphic, as anticipated, displaying values of PIC from 0.569 to 0.778.
Loci, alleles sizes and frequencies in Portuguese (53 wild vines and 57 cultivars) and foreign accessions
Loci, tamanho dos alelos e frequências nos acessos portugueses (53 plantas selvagens e 57 castas) e estrangeiros
In the fifty seven Portuguese native grapevine cultivars a total of fifty three alleles were scored across the six nuclear loci, thirteen alleles for locus VVS2 and eight for all the others. The most frequent allele was VVMD7-235 with a frequency of 0.465. The expected heterozygosity ranged from He=0.693 at locus VRZag79 to He=0.849 at locus VVMD5, with a mean value 0.767. With the exception of VrZag79 the observed heterozygosity was higher than the expected heterozygosity as a result of the absence of null alleles in all studied loci. All nuclear microsatellite loci scored were polymorphic, displaying values of PIC between 0.646 and 0.831.
When analyzing both Portuguese subspecies a total of sixty eight alleles were scored across the six nuclear loci (Table III). Fifty six percent of the alleles are present in both Portuguese subspecies. Seven alleles in wild vines and five in grapevine cultivars have frequencies above thirty percent. Both subspecies have twenty two percent of single alleles. In the wild vines the number of homozygosity by locus range from 26.4% (VVS2) to 49.05% (VVMD27) with an average of 40%. In the Portuguese grapevine cultivars this percentage is much lower, ranging from 5.26% (VVMD5) to 33% (VRZag79) with an average of 16.6%. In all loci, with the exception of locus VVMD5, there are alleles scored in the foreign cultivars also found at low frequencies in the Portuguese cultivars, but not found in Portuguese wild vines (Table III, underlined).
Chlorotype diversity
As for the portuguese cultivars (Cunha et al., 2009), in the nineteen international cultivars analyzed no polymorphisms were found for the ccmp2 locus. Using the designations proposed by Arroyo-Garcia et al. (2006) for the combination of the three polymorphic chloroplastidial loci (Supplementary Table I) the following chlorotype (chl) distributionwas found in international cultivars analysed: 10 cultivars belonged to chl A, 1 cultivar to chlB; 1 cultivar to chl C; and 7 cultivars belong to chl D. As in the case of the native Portuguese germplasm chl A is the most frequent (Cunha et al., 2009). The accession with chl B is Coarna Neagra, a native cultivar from Romania and the one with chl C is Ahmer Bou Amer a native from North Africa.
Wild vine intra-population diversity and Hardy-Weinberg (HW) equilibrium
The total number of nuclear microsatellite alleles per population ranged from 24, in population 04, to 38, in population 01 (Table IV). The mean number of alleles per locus and per population ranged from 4.0, in population 04, to 6.3, in population 01. The allelic richness (number of alleles per locus independent of sample size) ranged from 3.5, in population 04, to 4.8, in population 01. The rare alleles (alleles present in less than 5% of the individuals per population) ranged from zero in population 05, to 13, in population 01. The number of private alleles present per wild vine population is one or two. The observed heterozygosity per population ranged from Ho=0.545 (population 02) to Ho=0.729 (population 05) and the expected heterozygosity per population ranged from He=0.545 (population 04) to He=0.665 (population 01). A significant deviation from HW equilibrium was found in the locus VVMD27 on populations 02 and 04; in the loci VVMD5 and VVS2 on population 01; and in the locus VVS2 of population 01 (Table V - in bold). No consistent patterns across loci and populations were observed.
TABLE IV.
Population, sample size and genetic variability estimates based on data from six microsatellite loci in four wild vines population
População, tamanho da amostra e estimadores da variabilidade genética baseados na análise de seis loci de microsatélites nucleares em quatro populações de vinha selvagem
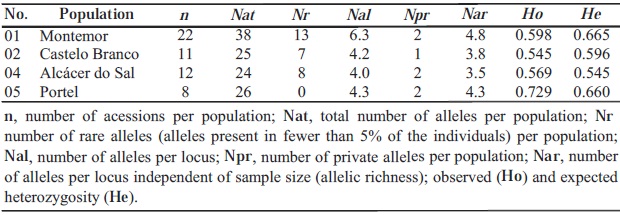
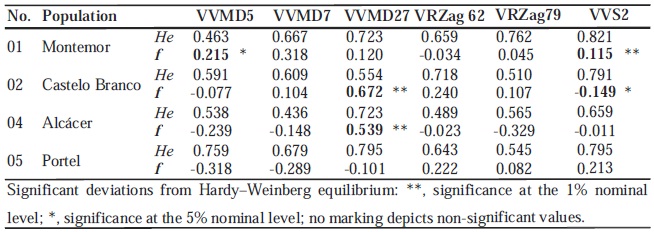
TABLE V.
Expected heterozygosity (He) and Inbreeding coefficient f (Weir and Cockerham, 1984) across six nuclear microsatellite loci in the four Portuguese wild vine populations.
Heterozigocidade esperada (He) e coeficiente de consanguinidade f (Weir e Cockerham, 1984) detectados com os seis loci de microsatélites nucleares nas quatro populações portuguesas de vinha selvagem.
Differentiation and genetic relationships among native Portuguese subspecies
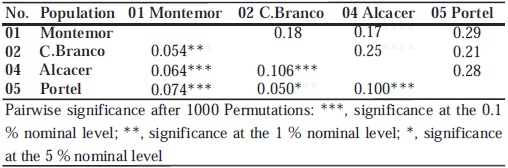
The Neis standard genetic distance and the pair wise genetic differentiation values (FST) among the four sylvestris populations are shown in Table VI. The higher FST values observed are between population 04 and populations 02 and 05, respectively 0.106 and 0.101. A lower but still significant value of FST is observed between the populations 05 and 02. A similar pattern of differentiation among populations is seen when calculating the Neis genetic distance (Table VI). The Analysis of Molecular Variance (AMOVA) showed that most of the genetic diversity was attributable to differences among individuals within populations (93.0%) but f values among populations are still significant (fst = 0.071; P, 0.001), showing a low inter-population differentiation.
TABLE VI.
Neis standard genetic distance (upper diagonal) and pairwise FST values (lower diagonal) among four wild vine populations
Distancia standardizada de Nei (diagonal superior) e valores de FST (diagonal inferior) entre as quatro populações de vinha selvagem.
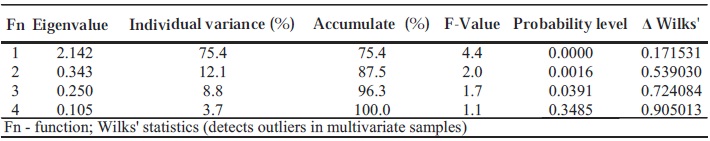
A multiple discriminant analysis was done to evaluate if the plants belonging to the different populations were correctly assigned. The first dimension explain 75.4% of the total polymorphism (Table VII), giving a good confidence to the discrimination among the different populations studied. This is stressed by the high significance (α = 0.0000) of Wilks lambda statistics (0.171) for this dimension. This analysis showed that some accessions could be assigned to other subspecies that the one they belong to.
TABLE VII.
Discriminant analysis among 53 wild plants from four populations and 57 native grapevine cultivars
Tabela VII - Análise discriminante entre as 53 plantas das quatro populações selvagens e as 57 castas nativas de Portugal
When using a 80% value of probability as the cut-off for incorrect assignment between the two subspecies some accessions could be identified (Supplementary table II): accession 0125, a male plant from population 01, and accession 0411, a female plant from population 04, have a probability of 81% and 87.3% respectively to belong to the viniferasubspecies; Espadeiro Mole, Trincadeira das Pratas, Antão Vaz, Monvedro and Cornifesto are attributable to the sylvestris subspecies by 98.1%, 93.0%, 87.3%, 86.2%, 81%, respectively.
A Principal Coordinate Analysis was used (with a minimum spanning tree superimposed) to find the relationships between all the genotypes studied (native Portuguese accessions from both subspecies and foreign grapevine cultivars) combining nuclear and chloroplastidial microsatellite data (Figure 2). The first two axes describe 50.4% of the total variance (31.3% and 19.1% respectively). The inclusion of chloroplastidial data allowed the separation of the accessions by maternal inheritance. Along the two axes, four groups are formed according to chlorotype. Chl A and B are subdivided being most of vinifera accessions clustered together and most of the sylvestris accessions forming sub-clusters that establish connections between both chlorotypes through accessions of the same populations. Some grapevine cultivars (Espadeiro Mole and Encruzado) are solely link to wild vines plants. Wild vine 0125 is tied directly to Uva Cão (figure 2, arrow). The wild vine 0107, a female plant, establishes the connection between wild vine population 01 and the larger group of grapevine cultivars.
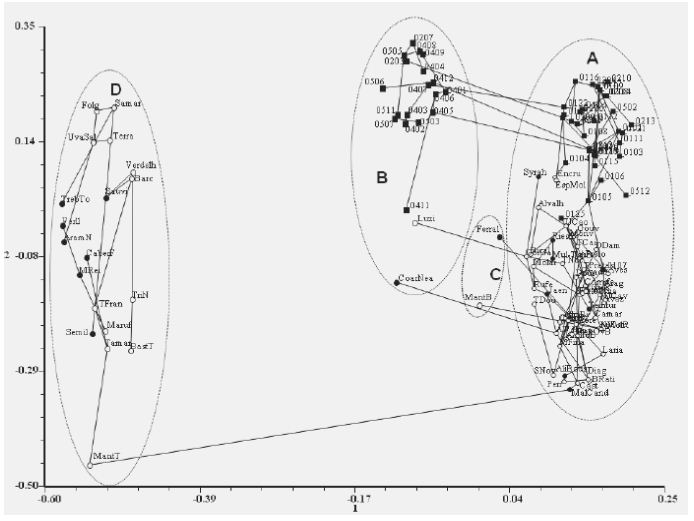
Figure 2 - Principal Coordinate Analysis of nuclear and chloroplastidial microsatellites data scored for Portuguese wild vines and Portuguese cultivated and foreign cultivated accessions. Minimum spanning tree superimposed. Portuguese wild vines – numbers; Portuguese and foreign grapevine cultivars – acronym; Portuguese accessions – open circles; foreign accession – closed circles. A, B, C and D - Chlorotypes.
Figura 2 – Análise de componentes principais dos microsatélies nucleares e cloroplastidiais das videiras selvagens e das castas portuguesas e estrangeiras em estudo. Sobreposição da árvore de expansão mínima. Videiras selvagens portuguesas – numeradas; Castas cultivadas portuguesas e estrangeiras – acrónimos; Acessos portugueses – círculos abertos. Acesos estrangeiros – círculos fechados. A, B, C e D - Clorotipos.
DISCUSSION
Using morphological traits as a sample strategy to differentiate between individuals revelled to be an efficient method to obtain a collection of wild vine plants, since only seven were found to be clones. Most of these clones were collected in the population 05 (Portel) were the major vine tutor were Robus sp. that difficult the observation of morphological traits. The use of the six recommended nuclear loci was sufficient to differentiate all genotypes. The large number of alleles scored (Table II) was anticipated from the observed morphological variation in native Portuguese Vitis vinifera (Cunha et al. 2009). An overall high genetic diversity is revealed across the native Portuguese accessions studied from both Vitis vinifera subspecies. The observed and expected heterozygosity values in wild vines (Ho = 0.601, He = 0.676) are lower than those of native grapevine cultivars (Ho = 0.833 and He = 0.767). These differences are probably due to introgression with foreign germplasm in the cultivated grapes, together with the severe selection strain exercised by introduced diseases in the nineteen century and the human pressure in the wild populations. Nevertheless the HW equilibrium observed at most loci and in most of the populations indicates an overall retained diversity in the existing wild vine populations. The high level of allelic diversity in these populations is most probably related to the mating system of dioecious and out-breeding plants.
Positive values of inbreeding coefficient (f)inwild vines populations might be due to geographic isolation that result in the emergence of homozygotic alleles and allele dropout. Loci VVMD5, VVMD27 and VVS2 show a relatively high deficit of heterozygosity in 01, 02 and 04 populations, possibly due to allele dropout.
To optimize the exploitation of natural diversity in wild vine it is important to identify the rare alleles, enabling the establishment of core collections for conservation and the study of the genetic evolution of this subspecies, as suggested by Le Cunff et al. (2008). The low allelic richness within wild vine populations 02 and 04 provides evidence for a potential genetic bottleneck effect caused by the stresses already referred. Allelic richness, being more heavily influenced by rare alleles than expected heterozygosity, is commonly regarded as more relevant criteria for measuring loss of diversity due to genetic bottlenecks (Nei et al., 1975).
The genetic diversity found in the native Portuguese vinifera subspecies is consistent with the one from other Portuguese vinifera samples (Lopes et al., 2006, Almadanim et al., 2007). When Le Cunff et al. (2008) determine the optimal size of a core collection to represent the available germplasm diversity of worldwide cultivated V. vinifera, two Portuguese grapevine cultivars are included in a core of twelve; two more in a core of forty eight and another two in a core of ninety two, this last collection representing 100% of total SSR diversity. The inclusion of these Portuguese cultivars in these core collections stresses the importance of the Portuguese gene pool in the overall cultivated grapevine gene pool.
Introgression with foreign gene pools in the cultivated Portuguese cultivated grapevines (Table III) is reinforced by the presence of alleles scored at low frequencies that are absent in the Portuguese wild vine populations and present in the foreign accessions studied. Also the Portuguese wild vines do not contain chlorotype C and D. Assuming that the origin of Portuguese cultivars was a result of the domestication of autochthonous germplasm, the presence of chlorotype C and D is a likely outcome of crosses among foreign introduced cultivars and local germplasm. An exclusive local domestication of grapevine will have restricted the chlorotypes to A and B in cultivated accessions. It is not obviously the case since 21% of the native grapevine cultivars have chlorotypes C and D (respectively 3.5% and 17.5%) attesting the contribution of foreign genotypes to the Portuguese germplasm. Taken together, the chlorotype analysis of the foreign and of the autochthonous germplasm reinforce the suggestion made by Arroyo-Garcia et al. (2006) that the western Mediterranean region was a centre of origin of the cultivated grapevines.
The AMOVA carried out in the Portuguese wild vines populations showed a high level of intra-population diversity and a low but still significant genetic differentiation among populations. Most of the genetic diversity was conserved inside each population as is the case for woody perennial out-breeding species that maintain most of their variation within populations (Belaj et al., 2007). No relation was found between geographic area and population genetic diversity.
Multiple discriminant analysis performed with nuclear microsatellites data from both subspecies showed intermingle within wild vines populations and among these and grapevine cultivars. Wild vines appear to form a continuum and there is no clear population division, reinforcing the idea that until the recent biotic stresses and human intervention all the populations were connected. The probability of misclassification of accession 0125, a male wild vine, may be attributed, taking into account the genetic determinism of sex in Vitis vinifera (Marguerit et al., 2009), either to a mutation on a feral plant or the result of a cross between a grapevine hermaphrodite cultivar and a male wild. Theoretically wild male plants cannot be escapes from the vinifera subspecies (either hermaphrodite or female cultivars) and cannot result from pollination between wild females and cultivated hermaphrodite or female plants. Accession 0411, a feminine wild vine plant classifiable as belonging to the vinifera subspecies, is probably derived from a cross between both subspecies.
The five grapevine cultivars attributable to the subspecies sylvestrismay be seen as prove that they were locally domesticated.
The assignment of accessions of one subspecies to the other one is also an indication of gene flow among both subspecies which is also indirectly observed by the presence of a Grapevine Ruspestris stem pitting virus variants, transmitted by pollen, in a number of infected female plants from wild vine populations imbedded in a grapevine growing area (Nolasco et al., 2006). It is notable that in the wild vine populations studied male plants were not infected with this virus.
Principal coordinate analysis using nuclear and chloroplastidial data reinforce some of the results obtained with multiple discriminant analysis using nuclear microsatellites. Wild accession 0125, a male wild plant assigned to the cultivated subspecies in the multiple discriminant analysis, is directly link to cultivar Uva Cão (Fig 2, arrow) in the principal coordinate analyses, stressing the probability that this accession could be a descendent of a feral form. Espadeiro Mole, a cultivar assigned to a wild population in the multiple discriminant analysis, is directly link to the wild accession 0206, stressing the probability that this accession could be a descendent from plants of the wild subspecies.
CONCLUSION
This study establishes the range of existing variability in the Portuguese native vine germplasm. It also provides a baseline for future monitoring of Portuguese vine genetic diversity. Intra-population genetic variation was similar across the geographic range, regardless of population size. While there seem not to be an immediate danger of genetic erosion in neither of the subspecies, some precautions should be taken: the number of wild populations is low and there is potential risk of disappearance due to river bank cleanings; the number of used cultivars is very low compared to the number of autochthonous available cultivars, and the loss of interest in their use may lead to their rapid disappearance.
This study shows that the autochthonous grapevine cultivars probably derive from local wild germplasm. This corroborates previous conclusions that the Iberian Peninsula has been a secondary centre for grapevine domestication. Nevertheless it was also verified that a certain amount of introgression from foreign germplasm can be found in some of the Portuguese cultivated germplasm.
The genetic richness of both wild and cultivated subspecies is a potential useful tool for breeding purposes. Maintenance of allelic richness especially rare alleles must be prioritized because these genotypes may harbour rare characteristics being potentially useful for breeding purposes or for the identification of molecular markers associated to particular environmental adaptations as well as some resistances to crop diseases. To prevent the loss of variability in the wild vine germplasm, we propose the development of an in situ program of conservation, together with a re-population of similar riparian wood environments. The implementation of a legal protected status for the subspecies sylvestris will be fundamental to guaranty the implementation of a meaningful conservation strategy. A continuing effort is being made to maintain collections of autochthonous cultivated grapevines. This effort must be sustained and amplified.
ACKNOWLEDGEMENTS
The authors thank the supported by FCT -PARIPIPI-Project A, GrapeGen06 EU project and Grasp Grape Wine EU project. Jorge Cunha was supported by a PhD scholarship (SFRH / BD /16226 / 2004) from Portuguese Ministry of Science Technology and Advanced studies.
REFERENCES
Almadanim M.C., Baleiras-Couto M.M., Pereira H.S., Carneiro L.C., Fevereiro P., Eiras-Dias J.E., Morais L., Viegas W., Veloso M.M.. 2007. Genetic diversity of the grapevine (Vitis vinifera L.) varieties most utilized for wine production in Portugal. Vitis, 46, 116-119. [ Links ]
Aradhya M.K., Dangl G.S., Prins B.H., Boursiquot J.M., Walker M.A., Meredith C.P., Simon C.J.. 2003. Genetic structure and differentiation in cultivated grape, Vitis vinifera L.. Genet Res Camb., 81, 179–192.
Arroyo-García R., Lefort F., de Andrés M.T., Ibáñaez J., Borrego J., Jouve N., Cabello F., Martínez-Zapater J.M. 2002. Chloroplast microsatellites polymorphisms in Vitis species. Genome , 45, 1142–1149.
Arroyo-García R., Ruiz-García L., Bolling L., Ocete R., López M.A., Arnold C., Ergul A., Söylemezolu G., Uzun H.I., Cabello F., Ibáñez J., Aradhya M.K., Atanassov A., Atanassov I., Balint S., Cenis J.L., Costantini L., Gorislavets S., Grando M.S., Klein B.Y., Mcgovern P.E., Merdinoglu D., Pejic I., Pelsy F., Primikirios N., Risovannaya V., Roubelakis-Angelakis K.A., Snoussi H., Sotiri P., Tamhankar S., This P., Troshin L., Malpica J.M., Lefort F., Martinez-Zapater J.M.. 2006. Multiple origins of cultivated grapevine (Vitis vinifera L. ssp sativa) based on chloroplast DNA polymorphisms. Mol. Ecol., 15, 3707–3714.
Belaj A., Muñoz-Diez C., Baldoni L., Porceddu A., Barranco D., Satovic Z. 2007. Genetic Diversity and Population Structure of Wild Olives from the North-western Mediterranean Assessed by SSR Markers. Annals of Botany, 1–10.
Buxó R.. 2008. The agricultural consequences of colonial contacts on the Iberian Peninsula in the first millennium B.C.. Veget. Hist. Archaeobot., 17, 145–154.
Crespan M.. 2004. Evidence on the evolution of polymorphism of microsatellite markers in varieties of Vitis vinifera L. Theor. Appl. Genet., 108, 231-237.
Cunha J., Teixeira Santos M., Carneiro L.C., Fevereiro P., Eiras-Dias J.E.. 2009. Traditional grapevine cultivars and wild vines (Vitis vinifera L.) are related in Portugal. Genet. Resour. Crop Evol. Doi., 10.1007/s10722-009-9416-4.
Doligez A., Bouquet A., Danglot Y., Lahogue F., Riaz S., Meredith C.P., Edwards K.J., This P.. 2002. Genetic mapping of grapevine (Vitis vinifera L.) applied to the detection of QTLs for seedlessness and berry weight. Theor. Appl. Genet. 105, 780–795.
Goudet J.. 1995. FSTAT (vers. 1.2): a computer program to calculate F-statistics. Journal of Heredity, 86, 485–486.
Grassi F., Labra M., Imazio S., Ocete Rubio R., Failla O., Scienza A., Sala F.. 2006. Phylogeographical structure and conservation genetics of wild grapevine. Conservation Genetics, ,,,7, 837–845.
Hegi G.. 1925. Illustrierte Flora von Mitteleuropa. 5, T.1, T. 3. Hansen Verlag, München. Germany.
Heywood V., Zohary D.. 1991. A catalogue of the wild relatives of cultivated plants native to Europe. Flora Mediterranea, 5, 375–415. [ Links ]
Imazio S., Labra M., Grassi F., Scienza A., Failla O.. 2006. Chloroplast microsatellites to investigate the origin of grapevine. Genet. Resour. Crop Evol. 53, 1003–1011.
Jaillon O., Aury J.M., Noel B., et al.. 2007. The grapevine genome sequence suggests ancestral hexaploidization in major angiosperm phyla. Nature, 449, 463–467.
Le Cunff L., Fournier-Level A., Laucou V., Vezzulli S., Lacombe T., Adam-Blondon A.F., Boursiquot J.M., This P. 2008. Construction of nested genetic core collections to optimize the exploitation of natural diversity in Vitis vinifera L. subsp. sativa. BMC Plant Biology, 8, 31.
Liu J.. 2002. POWERMARKER – A powerful software for marker data analysis. Raleigh, NC: North Carolina State University, Bioinformatics Research Center (http://www.powermarker.net).
Lopes M.S., Santos M.R., Eiras Dias J.E., Mendonça D., Câmara Machado A.. 2006. Discrimination of Portuguese grapevines based on microsatellite markers. Journal of Biotechnology, 127, 34–44.
Lopes M.S., Sefc K.M., Eiras Dias J.E., Steinkellner H., Laimer da Câmara Machado M., Câmara Machado A.. 1999. The use of microsatellites for germplasm management in a Portuguese grapevine collection. Theor. Appl. Genet. 99, 733–739.
López Martínez M.A., Ocete R., Pérez Izquierdo M.A.. 2001. El impacto antrópico sobre las poblaciones de vid silvestre: características sanitário-agronómicas y aprovechamiento de este recurso. In: Vine and Wine in European Culture. II International Symposium of the History and Civilisation of Vine and the Wine, Porto-Lamego-Vila Real, Portugal, September 10–12th.
Marguerit E., Boury C., Manicki A., Donnart M., Butterlin G., Némorin A., Wiedemann-Merdinoglu S., Merdinoglu D., Ollat N., Decroocq S.. 2009. Genetic dissection of sex determinism, inflorescence morphology and downy mildew resistance in grapevine. Theor. Appl. Genet. DOI 10.1007/s00122-009-0979-4.
De Mattia F., Imazio S., Grassi F., Baneh H.D., Scienza A.,·Labra M.. 2008. Study of Genetic Relationships Between Wild and Domesticated Grapevine Distributed from Middle East Regions to European Countries. Rendiconti Lincei 19, 223 – 240. DOI: 10.1007/s12210-008-0016-6.
Meredith C.P., Bowers J.E., Riaz S., Handley V., Bandman E.B., Dangl G.S.. 1999. The Identity and Parentage of the Variety Known in California as Petite Sirah. Am. J. Enol. Vitic. 50 (3), 236-242.
Minch E., Ruiz-Linares A., Goldstein D., Feldman M., Cavalli-Sforza L.L.. 1997. MICROSAT: a computer program for calculating various statistics on microsatellite allele data, ver. 1.5d. Stanford, CA: Stanford University, available at: http://hpgl.stanford.edu/projects/microsat.
Nei M., Maruyama T., Chakraborty R. 1975. The bottleneck effect and genetic variability in populations. Evolution, 29, 1–10.
Nolasco G., Santos C., Petrovic N., Teixeira Santos M., Cortez I., Fonseca F., Boben J., Nazaré Pereira A.M., Sequeira O. 2006. Rupestris stem pitting associated virus isolates are composed by mixtures of genomic variants which share a highly conserved coat protein. Arch. Virol. 151, 83–96.
OIV.. 2007. Descriptor list for grapevine varieties and Vitis species, OIV, Paris, 2nd edition.
Peakall R., Smouse P.E.. 2006. GENALEX 6: genetic analysis in Excel. Population genetic software for teaching and research. Mol. Ecol.. Notes, 6, 288–295.
Raymond M., Rousset F.. 1995. GENEPOP (version 1.2) population genetics software for exact tests and ecumenicism. J. Hered., 86, 248–249.
Rivera D., Walker M.J.. 1989. A review of paleobotanical findings of early Vitis in the Mediterranean and on the origin of cultivated grape-vines, with special reference to new pointers to prehistoric explotation in the Western Mediterranean. Rev. of Paleobotany, 6, 205-237.
Rohlf J.F.. 2000. NTSYS – pc: Numerical taxonomy and multivariate analysis system. Version 2.1 Exeter Software. Setauket. NY.
Rousset F., Raymond M. 1995. Testing heterozygote excess and deficiency. Genetics, 140, 1413–1419.
Schneider A., Carra A., Akkak A., This P., Laucou V., Botta R.. 2001. Verifying synonymies between grape cultivars from France and northwestern Italy using molecular markers. Vitis, 40, 197-203.
Sefc K.M., Lopes M.S., Lefort F., Botta R., Roubelakis-Angelakis K.A., Ibanez J., Pejic I., Wagner H.W., Glössl J., Steinkellner H.. 2000. Microsatellite variability in grapevine cultivars from different European regions and evaluation of assignment testing to assess the geographic origin of cultivars. Theor. Appl. Genet., 100, 498-505.
Sefc K.M., Regner F., Turetschek E., Glössl J., Steinkellner H.. 1999. Identification of microsatellite sequences in Vitis riparia and their applicability for genotyping of different Vitis species. Genome, 42, 1–7.
This P., Jung A., Boccacci P., Borrego J., Botta R., Costantini L., Crespan M., Dangl G.S., Eisenheld C., Ferreira-Monteiro F., Grando S., Ibañez J., Lacombe T., Laucou V., Magalhães R., Meredith C.P., Milani N., Peterlunger E., Regner F., Zulini L., Maul E.. 2004. Development of a standard set of microsatellite reference alleles for identification of grape cultivars. Theor. Appl. Genet., 109, 1448–1458.
Vendramin G.G., Lelli L., Rossi P., Morgante M.. 1996. A set of primers for the amplification of 20 chloroplast microsatellites in Pinaceae. Mol. Ecol., 5, 595–598.
Wan Y., Schwaninger H., Dan L., Simon C.J., Wang Y., Puchau H.. 2008. The eco-geographic distribuition of wild grape germpasm in China. Vitis, 47, 77–80.
Weising K., Gardner R.C.. 1999. A set of conserved PCR primers for the analysis of simple sequence repeat polymorphisms in chloroplast genomes of dicotyledoneous angiosperms. Genome, 42, 9–19.
* Corresponding author (351 21 441 1277. Email: psalema@itqb.unl.pt)
(Manuscrito recebido em 09.07.10 . Aceite para publicação em 23.07.10)